In this blog post, we are going to walk you through the development of a new feature that was recently released in Dataiku 9: distributed hyperparameter search. We will start with a quick reminder of what hyperparameter search is, why it is a very computationally intensive and slow process, and what it means to distribute it.
We will explore how the Dataiku platform works internally and discuss the challenges we faced while implementing this feature. Then, we will explain the solution we retained. At the end of this article, you will get the big picture of how distributed hyperparameter search works in Dataiku.
Hyperparameter Search: What Is It Already?
All learning algorithms have a variety of tuning parameters that must be adjusted in order to obtain the best results. These parameters, usually referred to as hyperparameters (HP), are not learned from data and must be set before training.
For example, the random forest training algorithm aggregates multiple decision trees together in order to model the relationship between several input features and a target variable. The number of trees, as well as their depths, are HPs that must be chosen beforehand.
It’s worth noting that there is no “ideal value” that always works best. It all depends on data patterns and, consequently, the best HP values will always be different. The process of finding out the best combination of HPs is called HP search.
Different Search Strategies
There is no easy way to determine beforehand the best combination of HPs for a given dataset. For this reason we must explore several combinations (i.e., each combination being referred to as a “HP point”) from all the possible ones (i.e., the “HP space”), compare them, and pick the one that works best.
The traditional way of performing HP optimization is grid search. Grid search is simply an exhaustive search over all possible combinations of HPs. To continue with the previous example about a random forest, we typically choose a few discrete values to explore for each HP (e.g., number of trees = [10, 50, 100], trees’ depth = [3, 5, 8]) and try all combinations.
It is well known that grid search is not always the best strategy, especially when the space is continuous or too large to be explored entirely. If you have a dozen HPs, combinatorial explosion makes it highly impractical to do an exhaustive search.
Advanced search strategies like random search and Bayesian search were introduced in Dataiku 8. In essence:
- Random search simply draws random points from the HP space and stops after N iterations.
- Bayesian search works like random search, but it generates new points depending on the performance of the previous ones. The idea is to focus the exploration around points that seem to work best.
Evaluating a Single HP Point
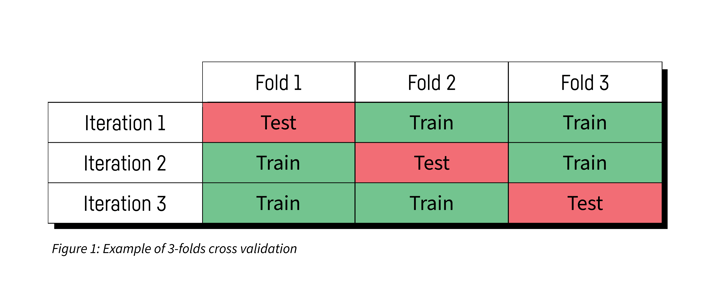
There exists several methods to evaluate an HP point, and the most commonly used is k-fold cross validation. The idea is to split the dataset into K folds and measure the model performance on each fold after training the model on the other K-1 folds. The measurements are then averaged over all folds in order to get the performance of the HP point.
Going into more details would be out of the scope of this article. The thing to remember here is that evaluating a single HP point requires training the model multiple times on different subsets of the dataset and that all these trainings are independent of each other.
An Embarrassingly Parallel Problem
The previous section should have given you an idea of what makes HP search a highly time consuming process.
From the previous, we can observe that:
1. A search strategy is a HP point generator.- Grid search and random search produce new points independently from the evaluation result of previous points.
- Bayesian search uses previous results to generate new points. We can’t know the whole set of points that need to be explored in advance. This also means that the level of achievable parallelism is limited.
2. The evaluation of a single HP point is constituted by k independent trainings (with k-fold cross validation).
HP search can be qualified as being an embarrassingly parallel problem. Indeed, it can be decomposed into many small and (almost) independent training tasks corresponding to each tuple (fold, HP point). This kind of problem is generally easy to distribute.
Distributed Search: How We Implemented It
Existing Implementation
Dataiku already had the ability to parallelize the HP search among multiple threads. Multithreaded HP search speeds up the process by the number of CPU cores. The implementation used by Dataiku is a fork of scikit-learn relying on the joblib library (see Figure 2).
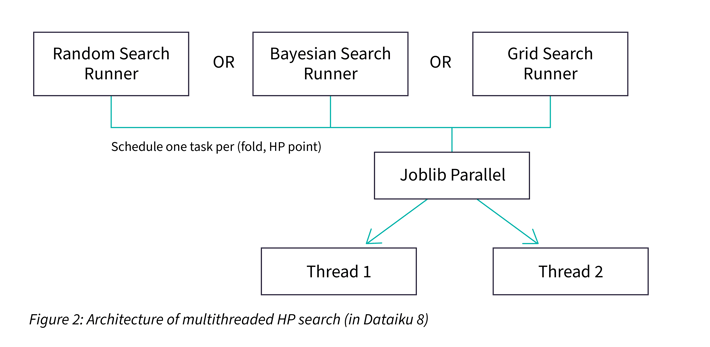
Notice that this implementation is limited by the capacity of a single machine, as it does not make sense to use more threads than CPU cores. Distributed search aims at leveraging the capacity of multiple machines in order to run the HP search faster.
Note that distributing the HP search on multiple machines only helps when the problem is "compute-constrained" (i.e., when the limiting factor is CPU time). This is typically the case when a lot of HPs need to be explored.
However, it cannot help when the problem is "memory-constrained" (when the dataset does not fit in memory) because the whole dataset is still required to fit into each machine’s memory. This second problem can only be addressed by using a distributed learning algorithm, which is a very different topic.
Helpful Context
Dataiku has a very good integration with Kubernetes clusters and can leverage them to push heavy computations out of the main server.
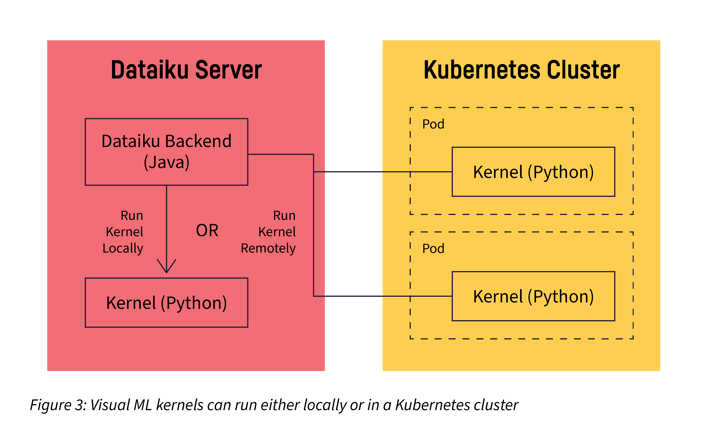
The main abstraction used by Dataiku is a “kernel.” A kernel is just a process, managed by Dataiku, which is running "somewhere." A kernel can be a local process or a process that runs remotely, typically in a Docker/Kubernetes container. The communication between Dataiku and the kernels is always network based.
In the context of visual machine learning (ML) in Dataiku, the training of a model is executed within a Python kernel in order to benefit from the excellent scikit-learn ML library of the Python ecosystem.
We initially questioned ourselves about reusing an existing distributed computing framework, versus rolling our own implementation.
We started experimenting with Dask but the results were not satisfying performance-wise. Moreover the integration of Dask within Dataiku was challenging because it introduced some capability overlap with what was already in place.
We finally decided to roll out our own implementation. Increased development and maintenance costs are compensated by having simpler integration with the Dataiku codebase and we also keep full control on all implementation details.
Development Plan
We considered two different approaches before starting the development of this feature:
- Create a PoC implementation as quickly as possible and iterate on it until it’s good enough.
- Plan ahead and build step by step, from the ground up.
From my experience, approach one is usually the best when there are unknowns or when the risk of failure is high, because it helps identify blockers or other unexpected problems earlier. Approach two is better when the scope is well defined and the goal is to ship something robust, because each module is built, properly tested, reviewed, and merged individually before they are all assembled to make the feature live.
In this case, we have chosen approach two because the main identified risk was to build something unreliable. Parallel code can be hard to get right and fixing bugs iteratively is often more costly than designing for correctness from the start.
Step 1: Dealing With Legacy Code
Dataiku’s HP search code is a several years old fork of scikit-learn and has since accumulated a lot of patches over time. It quickly became apparent that adding support for distributed search on top of it was not desirable because the end results would have been complex and hard to understand. Consequently, we decided to start the project by doing an iso-functional refactoring: rewrite the problematic code without changing its behavior.
The goals were:
- Explicitly introduce a worker interface which will later be compatible with remote workers (not yet implemented)
- The previous point implied the replacement of joblib by our own scheduler.
- Separate all the various concerns (strategies, k-fold, scheduler, worker API, etc.) in a way that makes them testable independently.
- Improve unit test coverage by taking advantage of this refactoring
The new structure looks like this:
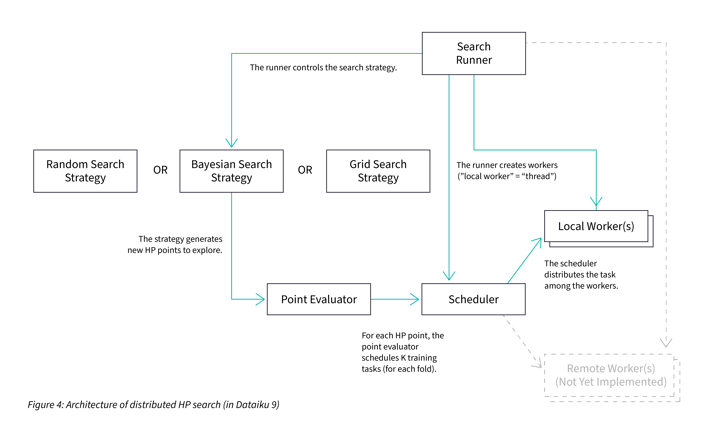
At a first glance, it looks like the new implementation (Figure 4) is more complex than the existing one (Figure 2). Moreover, the volume of code also increased by more than 50%. There is nothing wrong here: having more weakly connected boxes is actually a good thing — it shows a better separation of concerns. In fact, having only a few boxes was a sign that they were too large and hard to test.
Step 2: Kernel Communication
Having multiple machines cooperating on the same problem entails the need of communication between running kernels. How is it going to work?
As usual, we started looking at what was already in place. It turns out that Dataiku (written in Java) already had a protocol to communicate with its kernels (usually written in Python). We extended this protocol to allow direct “Python kernel to Python kernel” communication.
It is also worth noting that all networking questions were trivialized by the Kubernetes network model. It guarantees that all pods have an IP address in the cluster’s network and they can talk to each other regardless of their physical location (i.e., the node they are running on).
Finally, we need an efficient way to send Python objects over the network for different data structures. Cloudpickle was the simplest choice because it is able to serialize anything. Unfortunately, it had some known performance and robustness issues when dealing with large payloads — especially with old versions of Python — that we must support. At the end of the day, we ended up implementing a hybrid serialization scheme based on a mix of cloudpickle and custom serializers for large objects. It’s worth noting that Dask follows a similar hybrid approach, probably for the same reasons.
Step 3: Add Infrastructure to Spawn Remote Workers
The main infrastructure we needed was a mechanism in Dataiku that allows a kernel to request remote worker kernels whenever needed.
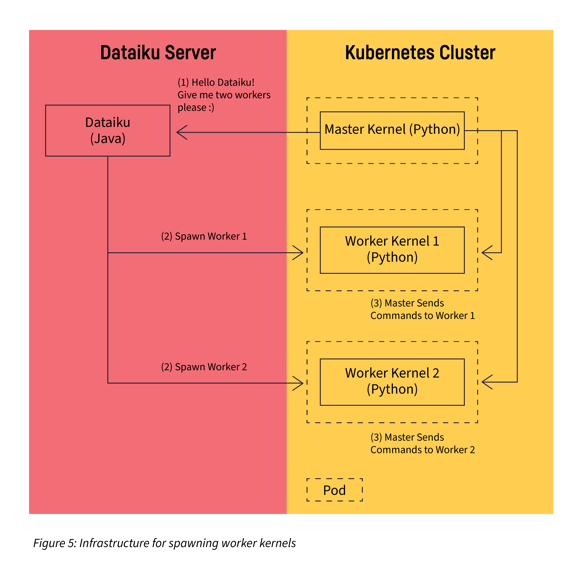
Step 4: Implementing Remote Workers
So, what do we have at this point?
- HP search code supports any worker as long as they implement the new worker interface.
- The master kernel is able to spawn remote worker kernels and talk to them.
The last module we need is an actual remote worker client implementation which:
- Conforms to the new worker interface, so that it can be used by the scheduler seamlessly
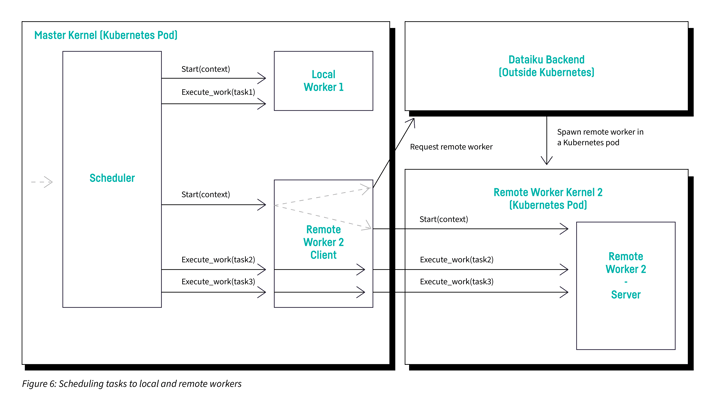
- Spawns a remote worker when it starts, transfers the data, and forwards the execution of tasks to the remote worker server
Figure 6 is a diagram showing how the client and server parts of the new remote worker interact together and with the scheduler. The remote worker client runs in the master kernel and the remote worker server runs in the remote worker kernel.
Benchmarks
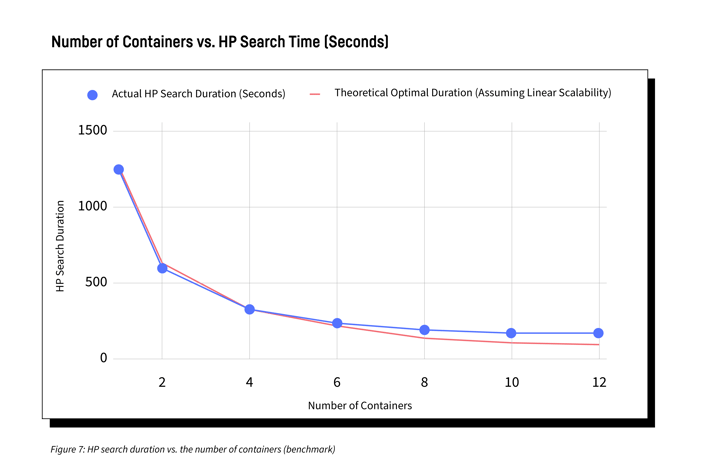
We performed some measurements in order to ensure the implementation was performing as expected. The following chart shows the time spent during HP search depending on the number of remote workers. As we can see, the performance increases almost linearly with the number of workers.
The difference with the “theoretical optimal duration” in Figure 7 becomes more visible when many workers are used. It comes from several factors:
- Remote container startup time
- Data serialization and network overhead
Conclusion
We’ve reached the end of our journey. I hope you enjoyed the ride and that you now have a better understanding of the kind of problems we tackle at Dataiku. I want to emphasize one last thing: this article was not really about implementing distributed HP search. What this article was really about is how we started implementing distributed computing capabilities natively into Dataiku. HP search was a perfect candidate for that due to its embarrassingly parallel nature, and it will also greatly benefit our users. This project opens the door to even more exciting projects!




