In this article, we try to reproduce the results from the 2019 paper “Diverse Mini-Batch Active Learning” and share our findings.
To begin, active learning is a process that automates data labeling through machine learning algorithms. By preparing data for labeling in an efficient way, models can perform better, increasing the business value of the output. Active learning methods typically aim to optimize one of the below criteria:
- Diversity. Diversity is the pure exploratory criterion. A method that purely optimizes for diversity would probably select samples all across the feature space and is more likely to select outliers.
- Representativeness. A sample is said to be representative if many samples are similar to it according to a given similarity measure. Typical representative samples are in the “center” of large sample clusters.
- Uncertainty. These are samples that are difficult for the model to classify.
Diversity and representativeness are usually optimized at the same time, given that both leverage the data distribution of unlabeled samples. Clustering methods optimize for both at once. This is the reason why the author of the paper has chosen the K-Means approach for his diverse query sampling method. Another strategy proposes a similarity-matrix-based method that integrates all criteria at once in a single loss function. Diversity is necessary to compensate for the lack of exploration of uncertainty methods, which mainly focus on regions close to the decision boundaries.
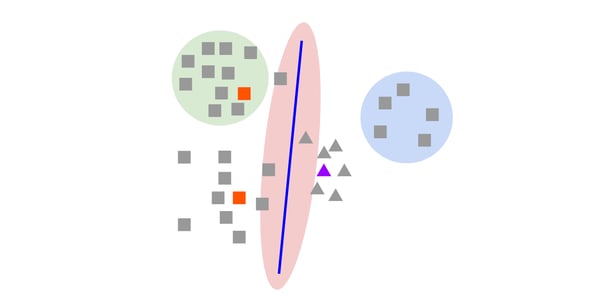 The above image shows an example of an active learning experiment on a binary classification task (square vs. triangle). The colored samples have been labeled and the current classifier boundary is shown in purple. Uncertainty-based methods are more likely to select samples in the red area, closer to the classifier boundary. Representative methods are more likely to select samples in the green area since the density of samples is higher. Only diversity-based methods are likely to explore the blue area. It shows that uncertainty or representativeness-based functions can miss the entire part of a feature space and fail to build a model with good generalization on unseen data.
The above image shows an example of an active learning experiment on a binary classification task (square vs. triangle). The colored samples have been labeled and the current classifier boundary is shown in purple. Uncertainty-based methods are more likely to select samples in the red area, closer to the classifier boundary. Representative methods are more likely to select samples in the green area since the density of samples is higher. Only diversity-based methods are likely to explore the blue area. It shows that uncertainty or representativeness-based functions can miss the entire part of a feature space and fail to build a model with good generalization on unseen data.
A natural way to segment the feature space is to use clustering methods. Since clusters are supposed to be composed of similar samples, a clustering method should both discriminate features and find representative samples within each cluster.
Inside Diverse Mini-Batch Active Learning
The diverse mini-batch active learning method combines uncertainty and diversity by selecting the next k samples to be labeled:
- First, pre-selecting β * k samples using the smallest margin sampler, β being the only parameter of the method.
- Then selecting k samples among this pre-selection by running either a submodular optimizer called Submodular(β), a K-Means called Clustered(β), or a K-Means weighted by the uncertainty score called WClustered(β). The paper reports experiments with β=10 and β=50.
The paper states that the method that solves the diversity problem best is submodular set function optimization and that K-Means is only a proxy to this problem. This subject is too complicated for a simple blog post, so we refer the curious reader to the original paper and the Wikipedia page! We have decided to compare both by using the submodular optimization from apricot python package and the K-Means optimization from scikit-learn.
Background for Choosing This Paper
There is no shortage of literature in the domain of active learning, but here are the top reasons we selected the “Diverse Mini-Batch Active Learning” paper:
1. A Sound Validation Scheme. We have noticed that a lot of active learning papers validate their method on small datasets coming from OpenML without enough samples to be considered in an active learning context — most often, 150 samples are enough to reach an accuracy of 0.95. We, therefore, wanted to reproduce the paper on a larger dataset. This experiment used CIFAR10, which seemed complex enough to be selected while staying reproducible without heavy resources.
2. A Simple Yet Elegant Method. Most recent papers focus on active learning for specific deep learning models; some of them are even dependent on the architecture of the model. We wanted to have a method that is simple enough to be understandable and reproducible. The proposed method is based on K-Means, a widely used clustering approach based on an intuitive principle.
3. Optimization for Both Uncertainty and Diversity. Most modern active learning methods optimize for both model uncertainty and feature exploration — also called diversity. The proposed method combines both of these criteria in a very natural way by using uncertainty as a sample pre-selection but also as a sample weight for the K-Means.
In the paper, the author compares its methods to a framework called FASS. For the sake of simplicity, we did not reproduce the results of this framework in this blog post. We have also decided to use Keras instead of MXNet out of habit and to see if findings can be reproduced using another framework.
Inside the CIFAR10 Experiment
This experiment is an image classification problem on the CIFAR10 dataset. Because bootstrapping this task requires a more extensive training set, the active learning loop starts with 1,000 samples, and adds batches of 1,000 samples until it reaches 10,000 samples (out of 50,000), and reaches an accuracy around 0.60. Our setting for this experiment was:
- We used Keras instead of MXNet and a Resnet 50 v2 instead of Resnet 34 v2 as it is not natively available in Keras. Since the paper specifies no optimizer, we chose to use RMSprop and ran 3 epochs.
- We used 50,000 samples for training and 10,000 for testing as in the paper.
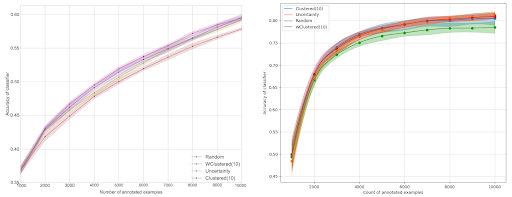
On the left, you can see the results of the original paper. On the right, you can see the results from our experiment on the CIFAR10 dataset. The performance measure is accuracy, the solid line is the average over 20 runs, and the confidence intervals are 10th and 90th percentiles.
So, what does this mean? The most striking result is that our accuracy is far higher than the one in the original paper, 0.8 instead of 0.6. The difference is likely due to the architectural difference between the two experiments. However, it is clear that active learning is better than random sampling even though we do not observe that the diversity-based sampling is more performant.
Conclusion
According to a 2019 survey from Cognilytica, data preparation and engineering tasks represent over 80% of the time consumed in most AI and machine learning projects. Active learning is one of those techniques that exists to help make the data labeling process more efficient, such as when not all data can be annotated because it is too costly or complicated or to optimize the order in which unlabeled data is processed.
While it can be difficult to reproduce experiments from a research article — particularly when not all of the parameters of the original experiment are available publicly — we are still pleased that we made the attempt. While our technologies and settings were slightly different, we were able to effectively reproduce the core findings and proved that active learning can be useful on complex labeling problems.




